This post and paper represents to me the huge problem, wasting hours of time and money, of using computers to make predictions and fancy images instead of doing something useful for humanity.
The paper ‘Infection-enhancing anti-SARS-CoV-2 antibodies recognize both the original Wuhan/D614G strain and Delta variants. A potential risk for mass vaccination?’ is based on modelling techniques.
What are these modelling techniques? Does the word make you feel a bit queasy, reminding you of Neil Fergusson’s modelling on how many people were gonna die? If so your gut instinct is correct.
The modelling used is described in the paper ‘Structural dynamics of SARS-CoV-2 variants: A health monitoring strategy for anticipating Covid-19 outbreaks’;-
‘In this study, we developed a global in silico analysis of virus binding’.
Whoa, hold your horses! They are not doing experiments, they are downloading data from each other, feeding it into a computer and then making some assumptions.
Here are some of the computations;
NONE of the proteins and gangliosides from the alleged variants in the pretty pictures have ever been shown to come from a virus (a transmissible and pathogenic entity);-
a) Electron microscopy of cell cultures shows virus like particles (these particles have never been seen in patient samples), some aspects of them, such as the ‘spike’ may be artefacts due to EM staining. These particles have never been shown to be viruses and are seen in uninfected cultures. The crystalline 3D structure of these virus like particles are observed using X-ray crystallography.
b) Some proteins, that have never been shown to be part of the virus like particles and are seen in healthy people, are chosen from crude samples or cell culture and their structure noted.
a) the 3D structure of the virus like particles not shown to be viruses is added to b) the structure of chosen proteins not shown to be part of the virus like particles and they are fed into a computer. The computer makes the chosen proteins fold and fit into the X-ray crystalline shape so that they form a 3D computer image. Hey presto! A 3D computer image of a thing with no basis in reality.
Here are some of the nonsense they conclude from their modelling in the paper:-
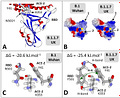
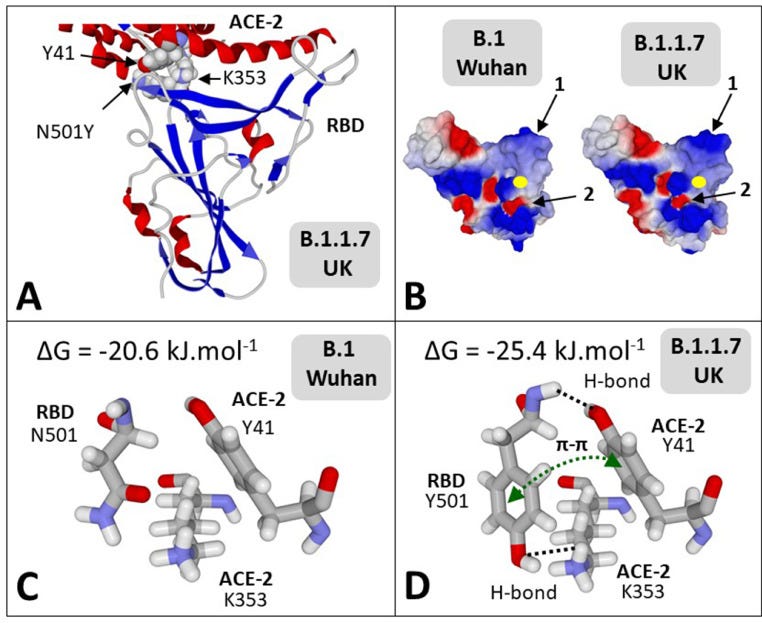
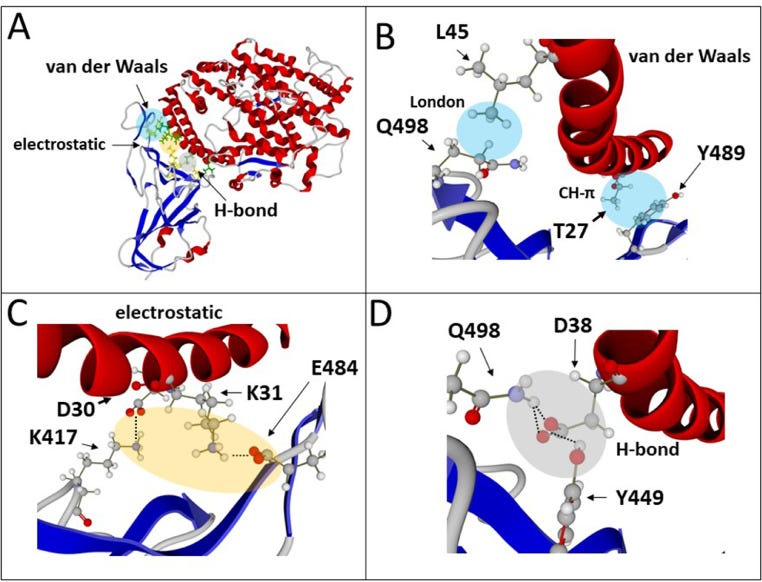
‘T-index takes into account the avidity of the NTD for lipid raft gangliosides (ΔGmut/ΔGwt [NTD-ganglioside]), the affinity of the RBD for ACE-2 (ΔGmut/ΔGwt [RBD-ACE-2]), and a global estimation of the electrostatic surface potential of both the NTD and the RBD (kinetic parameter). Values of T-index ranged from 2.16 for the Wuhan B.1 strain to 10.67 for the India B.1.617 variant (ca. 500% increase of transmissibility).’!!!
I think the best we can do is just ignore it.
Jo


Omg, I didn't even think an "in silico study" was a thing. People won't even know it was done on a computer. Pathetic.
By the way, I haven't been reading all the SK comments, but are you taking Steve Kirsch up on his challenge? Have you commented on this article?
https://stevekirsch.substack.com/p/if-viruses-dont-exist-then-how-can
thanks, good stuff. Appreciate the link to Christine Massey's work.
puzzling out a few things lately
Broxmeyer's 2011 Nexus article lays the foundation.
https://www.academia.edu/12968949/INFLUENZA_AND_THE_TUBERCULOSIS_CONNECTION_Part_1
Grant Genereux's hypothesis that they put Retinoic Acid in the inoculations is supported on a number of fronts; gleaned from his voracious scrubbing of papers for almost ten years now. If you want the full download over here let me know. Steve Kirsch ignored it back in November when he was all about sending vials to an independent lab.
Ran across this paper today, on a combo-search at PubMed - "Retinoic Acid and Tuberculosis".
https://pubmed.ncbi.nlm.nih.gov/34033876/
Retinoic acid induces antimicrobial peptides and cytokines leading to Mycobacterium tuberculosis elimination in airway epithelial cells
adds a bit of fuel to the fire